emgGO
Version 2.0 (4.75 MB) by
GallVp
A toolbox for offline muscle activity onset/offset detection in multi-channel EMG data.
emgGO
emgGO (electromyography, graphics and optimisation) is a toolbox for offline muscle activity onset/offset detection in multi-channel EMG data.
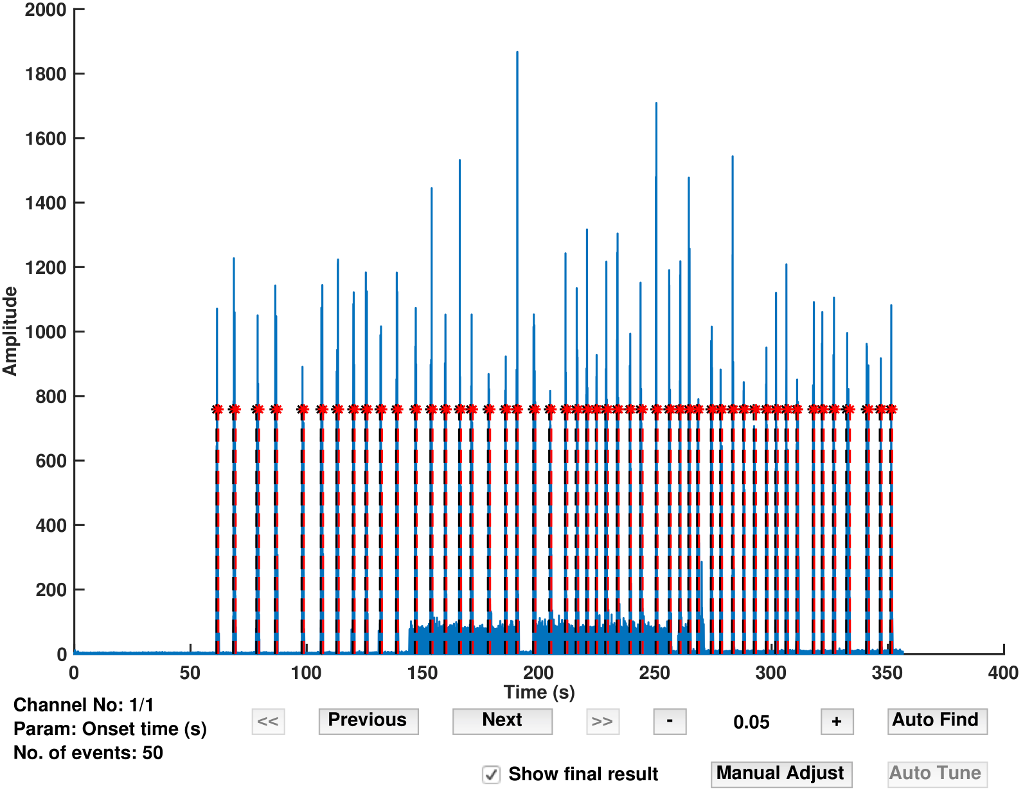
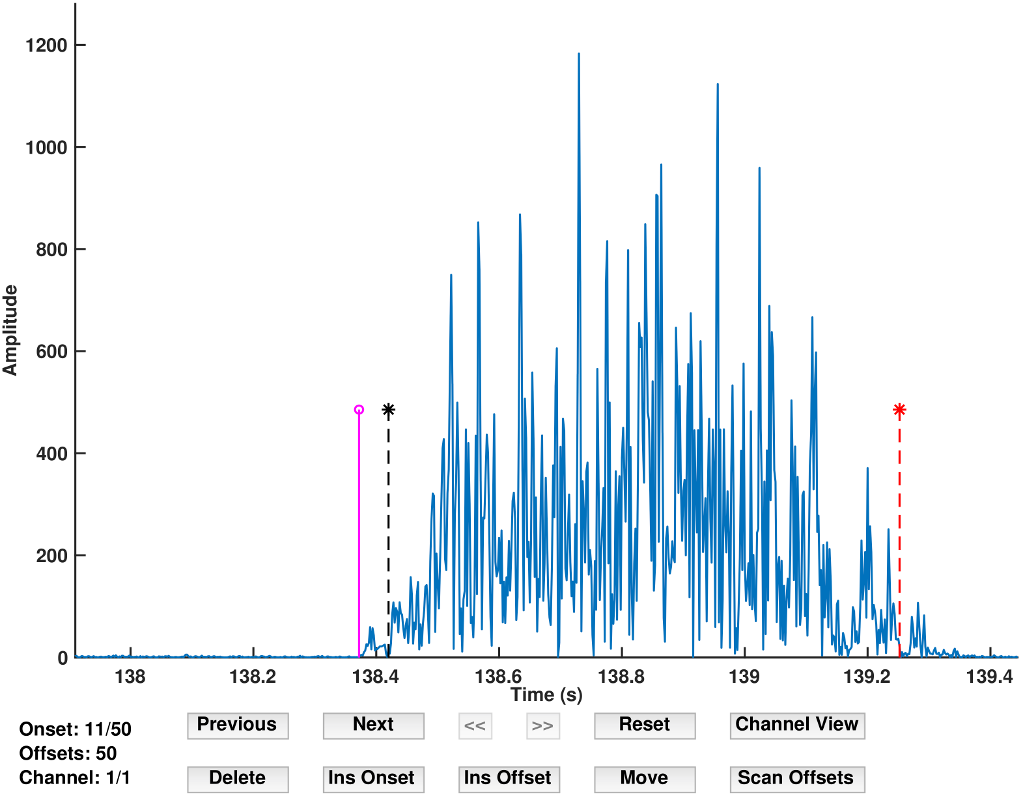
Fig 1. The GUI tools of emgGo which allow interactive processing of data.
Related Publications
- Optimal Automatic Detection of Muscle Activation Intervals, Journal of Electromyography and Kinesiology, doi: 10.1016/j.jelekin.2019.06.010
Compatibility
Currently emgGO is being developed on macOS Mojave, MATLAB 2017b.
Installation
- Clone the git repository using git. Or, download a compressed copy here.
$ git clone https://github.com/GallVp/emgGO
- From MATLAB file explorer, enter the emgGO folder by double clicking it. Follow the tutorials to experiment with the sample data.
Tutorials
- emgGO: An Overview
- How to Import Data in emgGO?
- How to Detect Onsets/Offsets?
- How to Create a Processing Pipeline?
- The Extended Double Thresholding Algorithm
Third Party Libraries
emgGO uses following third party libraries. The licenses for these libraries can be found next to source files in their respective libs/thirdpartlib folders.
Cite As
GallVp (2026). emgGO (https://github.com/GallVp/emgGO/releases/tag/v2.0), GitHub. Retrieved .
MATLAB Release Compatibility
Created with
R2017b
Compatible with any release
Platform Compatibility
Windows macOS LinuxTags
Discover Live Editor
Create scripts with code, output, and formatted text in a single executable document.
| Version | Published | Release Notes | |
|---|---|---|---|
| 2.0 |
To view or report issues in this GitHub add-on, visit the GitHub Repository.
To view or report issues in this GitHub add-on, visit the GitHub Repository.